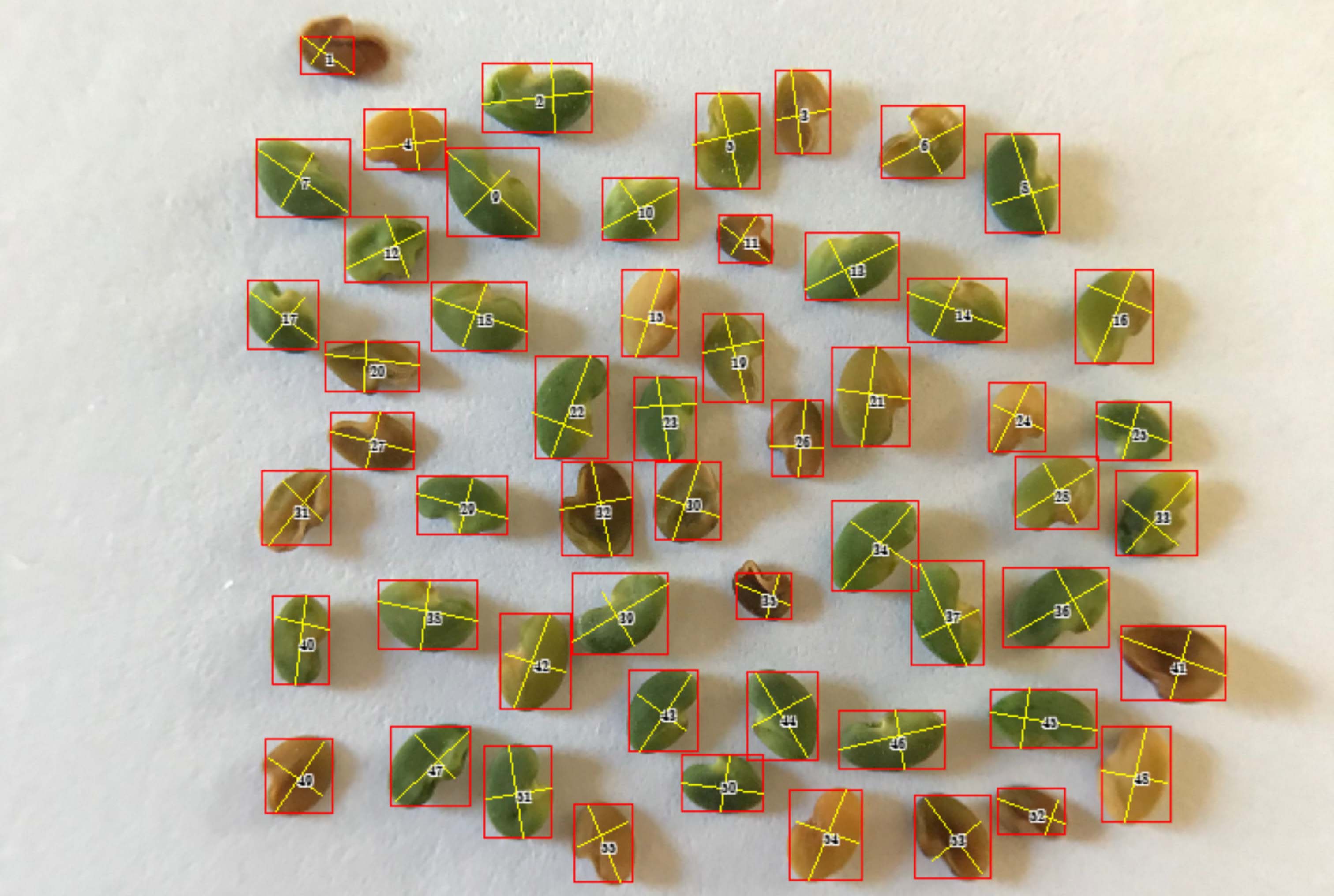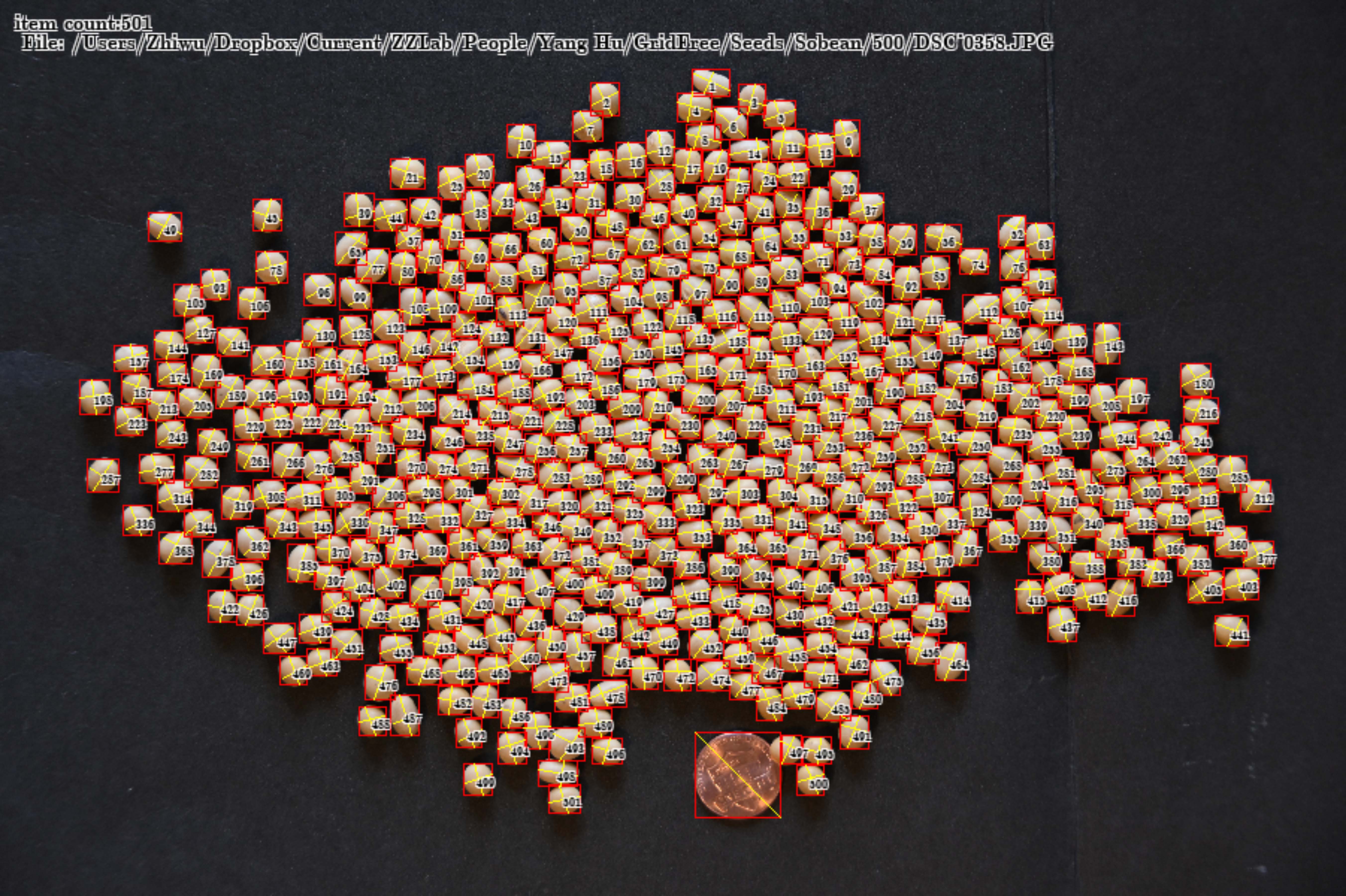
Grain characteristics are critical component traits for grain yield, including kernel length, kernel width, and thousand kernel weight. Manual measurements and counting are expensive, forming the bottleneck for the dissection of their genetic architecture toward the ultimate genetic improvement on grain yield. High throughput phenotyping methods have been developed by analyzing images of spreaded kernels on a surface. The challenges remain in segmentation of kernels from background variation, or separating kernels that are next to each other closely. In this study, we developed a software named GridFree to overcome these challenges. GridFree uses an unsupervised machine learning approach K-Means to separate kernels from background by using color index-based approaches, and uses Gaussian normal distribution as a dynamic criterion for a divide-and-combine strategy to segment adjacent kernels. When adjacent multiple kernels are incorrectly segmented as a single object, they form an outlier on the distribution of kernel areas, length, and width. GridFree allow users to choose an objects with multiple kernels as cutoff for splitting. In addition to counting, GridFree provides the measurement length, width, and area of kernels with option of either scaling with reference, not scaling. Benchmark evaluation against existing software demonstrated that GridFree had the smallest error on counting seeds of multiple crops, including alfalfa, canola, lentil, wheat, and soybean. GridFree was implemented in Python with a friendly graphic user interface. Users can fully stay in mental process with sliding bars, which completely liberate mechanical repetitive labor.




