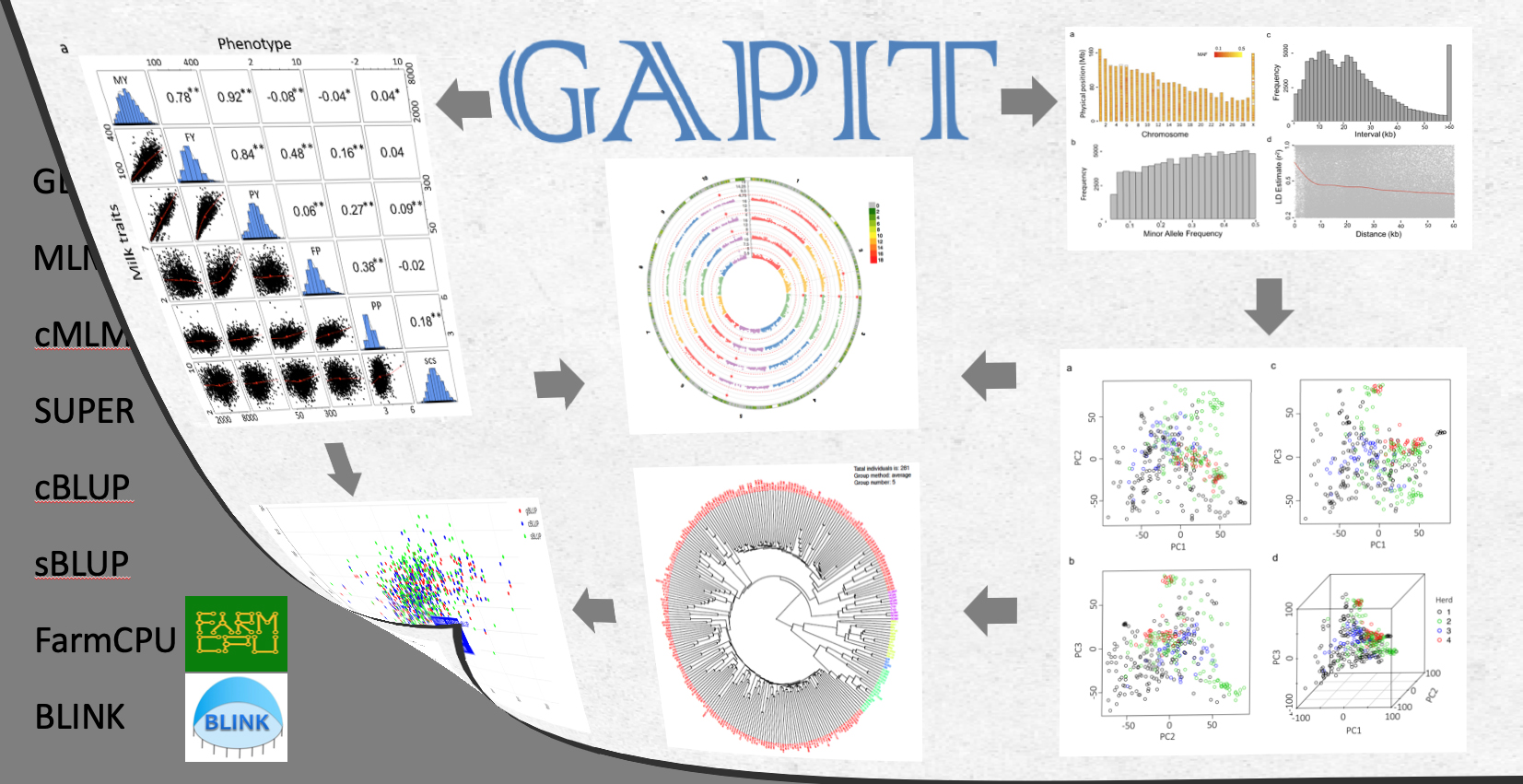
GAPIT is a Genome Association and Prediction Integrated Tool freely available for Public since 2011. It has been updated frequently to incorporate the state of art methods for Genome Wide Association Study (GWAS) and Genomic Selection (GS). The first two milestones of implementations were documented by the Bioinformatics paper led by Dr. Alex Lipka in 2012, and the Plant Genome paper led by Dr. You Tang and Dr. Xialolei Liu in 2016. Currently, Dr. Jiabo Wang is leading the new development, GAPIT version 3. In addition to the methods implemented in version 1 and 2 (e.g. Q+K, Compressed MLM, and SUPER), the new version implemented two new GWAS methods (FarmCPU and BLINK) and two new GS methods (sBLUP and cBLUP) published since 2016. All GAPIT documents are hosted at GitHub, including source code, user manual, demo data and demo code. The previous versions of source code are achieved at Dr. Zhiwu Zhang Lab (GAPIT achieve). GAPIT Forum is available for asking questions or giving comments. Please maximize the usage of the forum for benefit of our user community before contacting the current leading author (Dr. Jiabo Wang, email: wangjiaboyifeng@163.com).

